Research
Over a decade, the Keiser Lab developed AI and machine learning across a progression of scientific challenges: from predicting how drugs interact with protein targets, to decoding complex biological systems, to advancing the AI methods themselves.
Systems pharmacology & molecular prediction
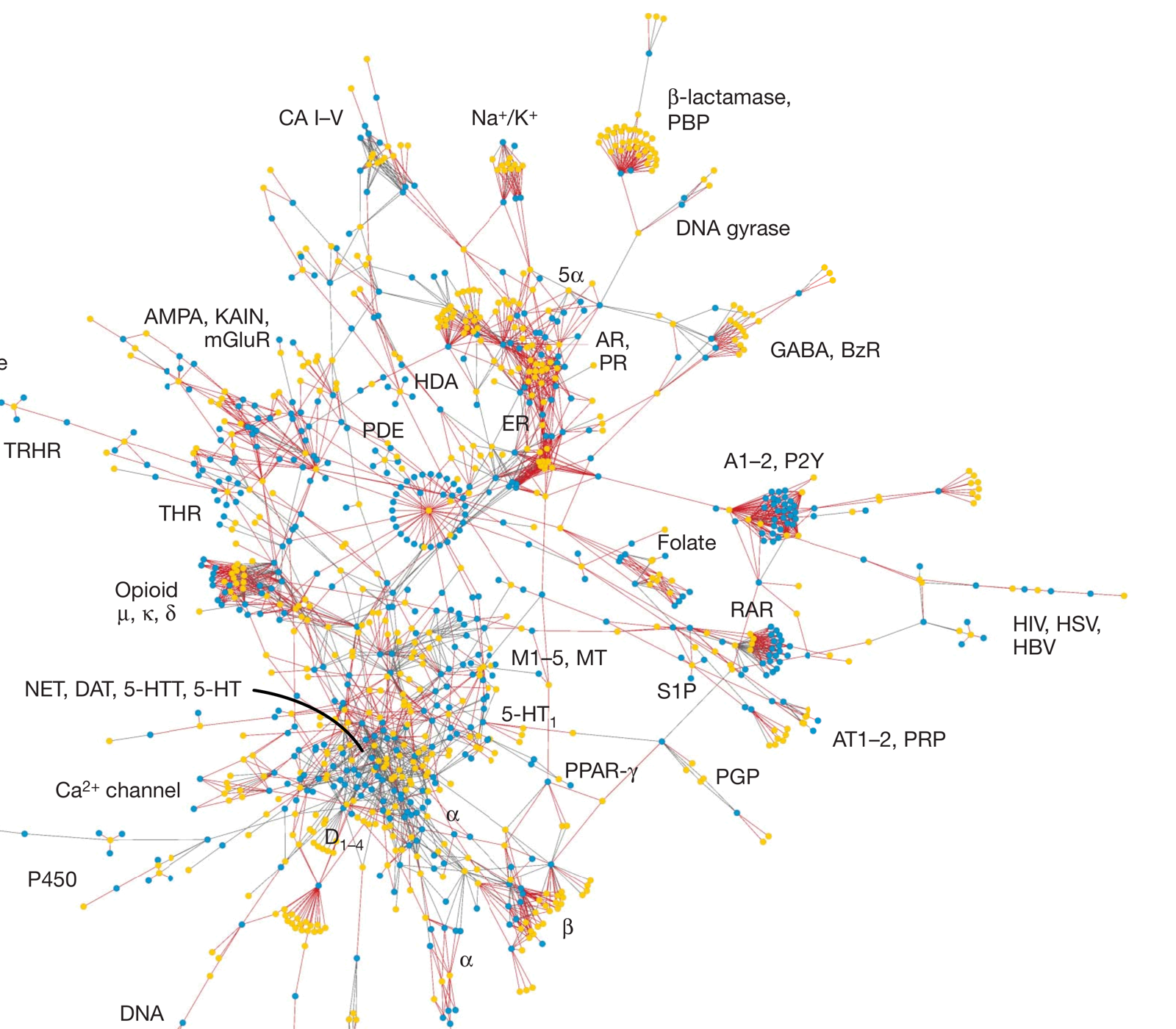
The lab’s earliest work addressed a foundational problem: drugs rarely act on a single target. The Similarity Ensemble Approach (SEA) used the statistical similarity of ligand sets to predict unexpected drug-target interactions, revealing that many approved drugs hit targets no one had anticipated (Nat Biotechnol 2007; Nature 2009). These predictions were validated at scale, identifying clinically relevant off-target effects across hundreds of drugs (Nature 2012) and informing FDA drug safety surveillance (Clin Pharmacol Ther 2015).
To move beyond ligand similarity, the lab developed new molecular representations using deep learning. E3FP encoded 3D molecular shape into learnable fingerprints (J Med Chem 2017), and later work advanced molecular representation learning for medicinal chemistry more broadly (J Med Chem 2020). These efforts laid the groundwork for increasingly powerful AI approaches to molecular interaction.
AI for complex biological systems
The lab applied deep learning to problems where the biological complexity exceeded what traditional computational methods could address.
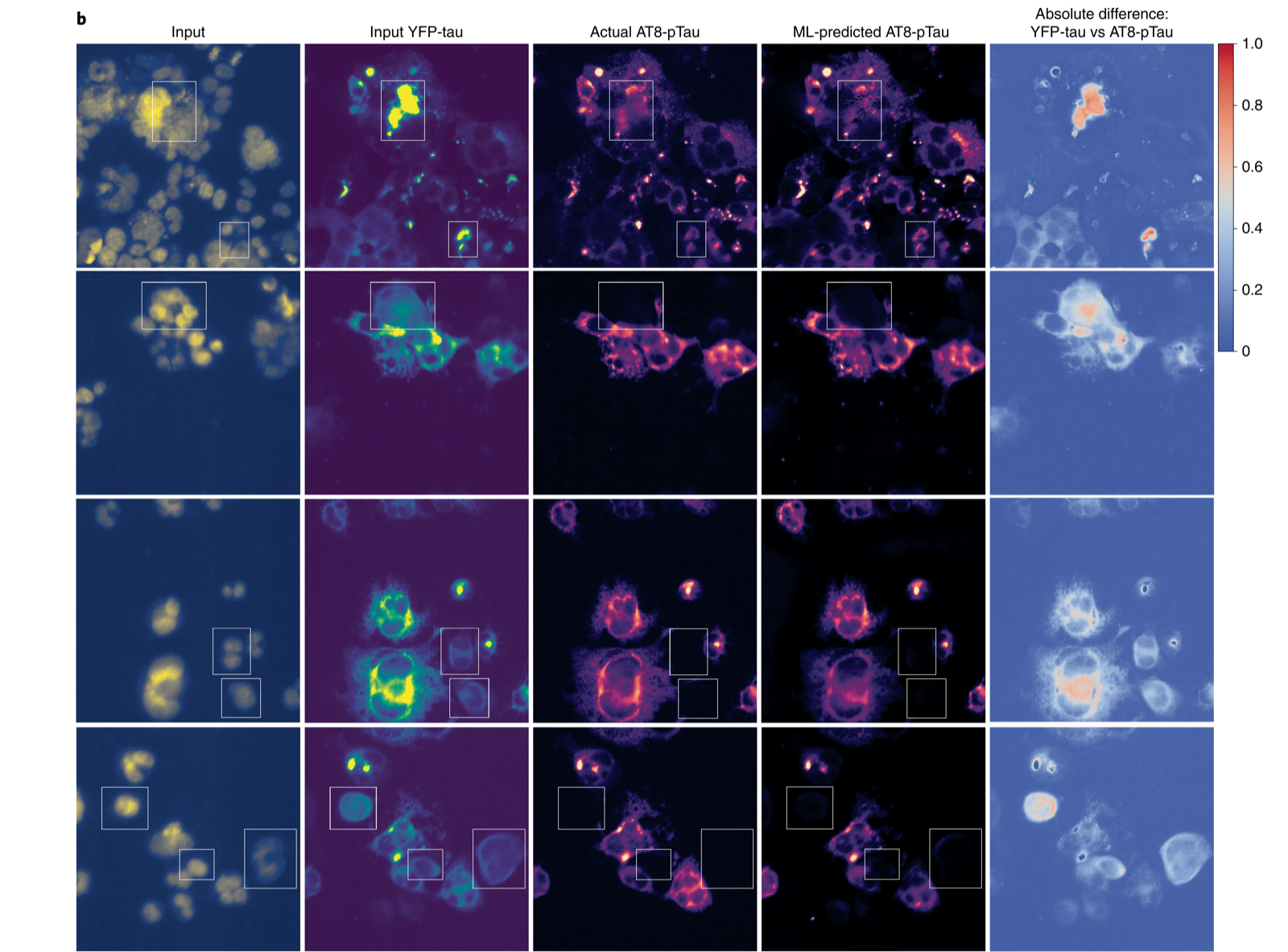
In phenotypic profiling, deep metric learning models decoded zebrafish behavioral responses to thousands of neuroactive compounds, identifying drugs with novel mechanisms of action that conventional target-based screening would miss (Nat Commun 2024; Nat Commun 2019). Trans-channel fluorescence learning generated informative image channels from high-content screens, unlocking historical datasets for Alzheimer’s drug discovery (Nat Mach Intell 2022).
In neuropathology, the lab built convolutional neural network pipelines for interpretable classification of Alzheimer’s disease pathologies (Nat Commun 2019), validated amyloid detection models across institutions (Acta Neuropathol Commun 2020; Commun Biol 2023), and developed tangle-tracer for precise neurofibrillary tangle segmentation from rapid point annotations. This work contributed to network-directed combination therapies for neurodegeneration (Cell 2025).
In genomics, the lab discovered that repetitive elements serve as key determinants of 3D genome folding (Cell Genom 2023) and developed ChromaFactor for deconvolution of single-molecule chromatin organization (PLoS Comput Biol 2025).
Advancing AI methods for science
The lab’s work progressively advanced the frontier of AI methods, not just applying existing models to scientific problems but developing new architectures, training paradigms, and evaluation frameworks.
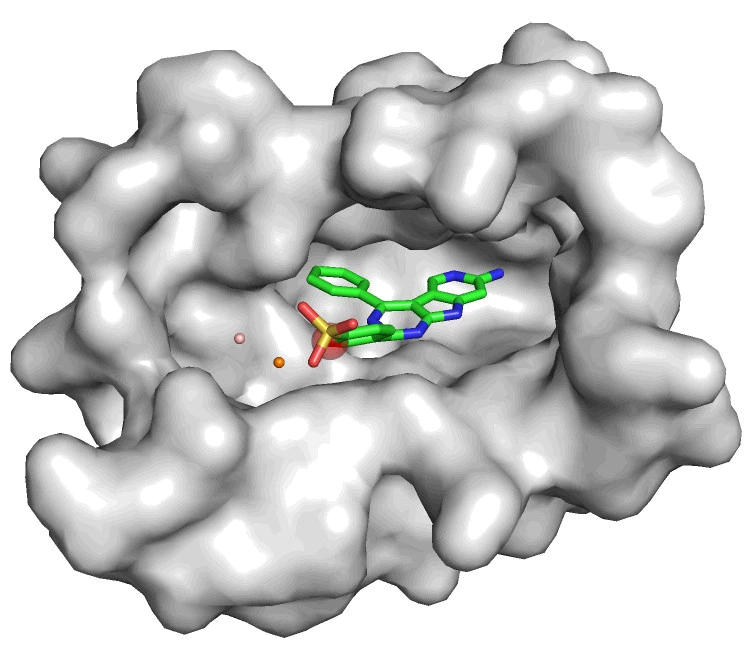
Generative models for molecular design represented the most forward-looking direction. AutoFragDiff introduced autoregressive fragment-based diffusion for pocket-aware ligand design, combining diffusion generative models with the structural constraints of protein binding sites (NeurIPS GenBio 2023). Attention-based learning on molecular ensembles explored how attention mechanisms could capture conformational diversity (NeurIPS ML4Molecules 2020). And the exceiver, a single-cell gene expression language model, applied transformer architectures to learn biological representations directly from transcriptomic data (NeurIPS LMRL 2022).
Rigorous evaluation of AI was a parallel theme. The lab demonstrated that adversarial controls are essential for scientific machine learning, showing how standard benchmarks can mislead (ACS Chem Biol 2018; Science 2018), developed robust concept activation vectors for semantic interpretability (ICML WHI 2020), and stress-tested diagnostic AI models for clinical readiness (NPJ Digit Med 2021). Methods for stochastic negative sampling improved molecular bioactivity prediction (J Chem Inf Model 2020), and VAE-based anomaly generation provided new tools for fuzz testing molecular representations (J Chem Inf Model 2025).
Retrieval-augmented and scalable approaches connected learned representations with efficient search. RAD applied hierarchical navigable small worlds to molecular docking (J Chem Inf Model 2024), and autoparty used machine learning to automate the visual inspection bottleneck in structure-based drug discovery (J Chem Inf Model 2025).